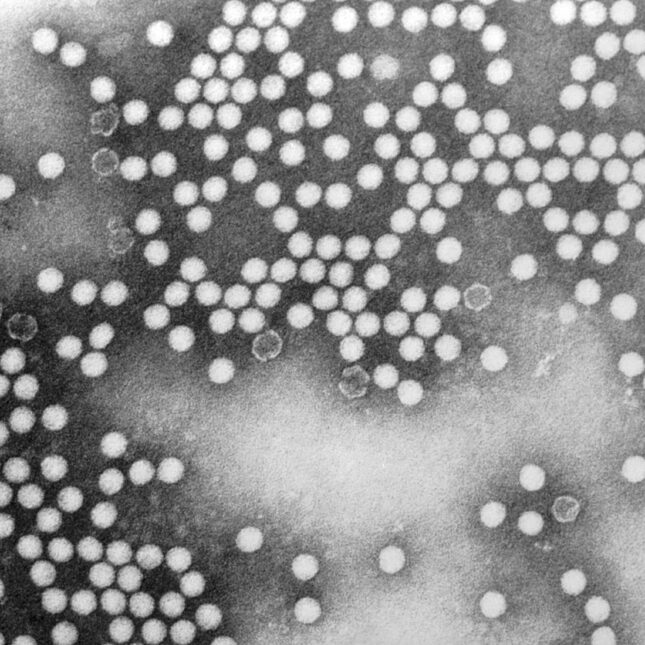
Researchers at the Earlham Institute have unveiled MARTi (Metagenomic Analysis in Real-Time), an innovative open-source software that enables real-time visualization of metagenomic data generated via nanopore sequencing. Published in Genome Research, this tool is poised to enhance the detection of microbial threats across clinical, agricultural, and environmental domains.
Originating from software designed for rapid pathogen identification in preterm infants, MARTi has been adapted for diverse applications, including use on research vessels in the Antarctic and in biosurveillance. The software comprises the MARTi Engine for analysis and a web-based GUI for result visualization, supporting multiple classification methods such as BLAST and Kraken2.
With antimicrobial resistance (AMR) posing significant challenges to global health, MARTi’s real-time updates on microbial community composition and AMR genes are particularly timely. Its integration with AirSeq technology allows for continuous air sampling and analysis, potentially transforming agricultural practices by providing farmers with immediate alerts on microbial threats. MARTi has demonstrated robust performance in both simulated and real-world datasets, marking a significant advancement in the accessibility and functionality of metagenomic analysis.
Open the full market picture for your next decision →